Fast microbubble dwell-time based ultrasonic molecular imaging approach for quantification and monitoring of angiogenesis in cancer
Abstract
Purpose: To develop and test a fast ultrasonic molecular imaging technique for quantification and monitoring of angiogenesis in cancer.
Materials and methods: A new software algorithm measuring the dwell time of contrast microbubbles in near real-time (henceforth, fast method) was developed and integrated in a clinical ultrasound system. In vivo quantification and monitoring of tumor angiogenesis during anti-VEGF antibody therapy was performed in human colon cancer xenografts in mice (n=20) using the new fast method following administration of vascular endothelial growth factor receptor 2 (VEGFR2)-targeted contrast microbubbles. Imaging results were compared with a traditional destruction/replenishment approach (henceforth, traditional method) in an intra-animal comparison.
Results: There was excellent correlation (R2=0.93; P<0.001) between the fast method and the traditional method in terms of VEGFR2-targeted in vivo ultrasonic molecular imaging with significantly higher (P=0.002) imaging signal in colon cancer xenografts using VEGFR2-targeted compared to control non-targeted contrast microbubbles. The new fast method was highly reproducible (ICC=0.87). Following anti-angiogenic therapy, ultrasonic molecular imaging signal decreased by an average of 41±10%, whereas imaging signal increased by an average of 54±8% in non-treated tumors over a 72-hour period. Decreased VEGFR2 expression levels following anti-VEGF therapy were confirmed on ex vivo immunofluorescent staining.
Conclusions: Fast ultrasonic molecular imaging based on dwell time microbubble signal measurements correlates well with the traditional measurement method, and allows reliable in vivo monitoring of anti-angiogenic therapy in human colon cancer xenografts. The improved work-flow afforded by the new quantification approach may facilitate clinical translation of ultrasonic molecular imaging.
Key words
Angiogenesis; cancer; ultrasound; microbubbles; molecular imaging; anti-VEGF therapy; dwell time
Introduction
Ultrasonic molecular imaging is increasingly being explored for cancer imaging (1-7), including early detection (8), molecular profiling (9,10), and therapeutic monitoring at the molecular level (11-14). For ultrasonic molecular imaging, gas-containing contrast microbubbles are functionalized by attaching ligands onto the microbubble shells that specifically bind to certain molecular markers under consideration (15). Contrast microbubbles remain exclusively within the vascular compartment following intravenous administration since their size of several micrometers prevents them from leaking into the extravascular space. Therefore, ultrasonic molecular imaging is particularly useful for imaging, monitoring and quantification of disease processes such as tumor angiogenesis that are characterized by molecular markers differentially expressed at the luminal site of endothelial cells of the tumor neovasculature (5,15).
After microbubble binding to the molecular target under consideration, in vivo ultrasonic molecular imaging signal is generated by a combination of backscattering owing to the difference of acoustic impedance of the microbubble gas compared to surrounding tissue and non-linear oscillations of microbubbles (16). At clinically used frequencies of several MHz, microbubbles resonate with non-linear oscillations which can be measured as harmonic or subharmonic frequencies of the imaging frequency, thereby enhancing signal to noise ratios from attached microbubbles compared to surrounding tissues that only show minimal non-linear properties beyond the second harmonic (15,17).
Most current preclinical ultrasonic molecular imaging protocols quantify molecularly-attached contrast microbubbles in vivo by using the traditional destruction/replenishment method (3,17): Several minutes following intravenous administration to allow binding of targeted microbubbles to the molecular target, a first ultrasound data set is acquired which reflects the sum of imaging signal from tissue signal, molecularly-attached microbubbles, and still freely circulating microbubbles. Following a high powered ultrasound pulse to destroy both attached and freely circulating microbubbles within the beam elevation of the ultrasound transducer, a second ultrasound data set is acquired at several seconds after the destruction pulse to allow freely circulating microbubbles to replenish into the imaging plane. The ultrasound imaging signal from microbubbles attached to the molecular target is then expressed as the difference of ultrasound imaging signal before and after the destruction pulse (18). While this approach is a robust technique and has been successfully used in preclinical in vivo ultrasound imaging with good correlation with ex vivo expression levels of molecular markers (9-14,19,20), it necessitates time-consuming post-processing, hampering the real-time work flow of ultrasound imaging. Additionally, ultrasound pressure necessary for microbubble destruction is not standardized for different microbubble types (21-23) and high powered destructive pulses for diagnostic purposes may cause unwarranted biological effects that are still not fully characterized (24,25). Therefore, a quasi real-time technique that allows automatic differentiation between imaging signal from attached versus freely circulating microbubbles without the need of a destructive pulse would be desirable. This will further facilitate clinical translation of ultrasonic molecular imaging and will allow integration of this technique into routine real-time ultrasound imaging protocols.
The purpose of our study was to develop and test a fast ultrasonic molecular imaging technique for quantification and monitoring of tumor angiogenesis during anti-angiogenic therapy and to compare this technique to the traditional destruction/replenishment method of ultrasonic molecular imaging signal quantification.
Material and methods
Human colon cancer xenograft model in mice
All experimental procedures using laboratory animals were approved by the Institutional Administrative Panel on Laboratory Animal Care. Human LS174T colon adenocarcinoma cells (ATCC; Manassas, VA) were cultured in Minimum Essential Medium supplemented with 10% fetal bovine serum to 70-80% confluency. Three million cells were trypsinized, centrifuged, and resuspended in 50 μL of matrigel (BD Biosciences, San Jose, CA) and then injected subcutaneously on the hindlimbs of 6-8 week old, female nude mice (n=20; Charles River, Wilmington, MA). Human colon cancer xenografts were allowed to grow for up to 7 days; tumor volumes were calculated using the formula for a prolate ellipsoid: π/6 × length × width × height, as measured by Brightness (B)-mode ultrasound.
Contrast microbubbles for contrast-enhanced ultrasound imaging
Two types of contrast microbubbles were used: control non-targeted perfluorobutane-containing microbubbles and clinical grade perfluorobutane-containing, lipopeptide-shelled microbubbles targeted to the human kinase insert domain receptor (which cross-react with mouse vascular endothelial growth factor receptor type 2, VEGFR2; mean diameter of 1.5 μm (range, 1-3 μm); BR55, Bracco Suisse SA, Geneva Switzerland) (14,26). Both control and VEGFR2-targeted contrast microbubbles (5×107 microbubbles in 180 μL volume per injection) were administered intravenously in random order to all mice through a tail vein catheter (Micromarker kitTM; VisualSonics, Toronto, Canada) at a constant injection rate of 1.2 mL/min (9 sec for 180 μL) using an infusion pump (Genie Plus; Kent Scientific, Torrington, CT). A waiting time of at least 30 minutes between contrast microbubble injections was applied to allow microbubbles from previous injections to clear (14,19); background levels were also measured to ensure microbubble clearance was achieved (through comparison with background levels prior to any injection of microbubbles).
Ultrasound imaging system
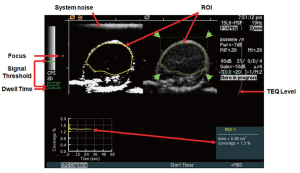
All human colon cancer xenografts were imaged with a Siemens Sequoia Acuson 512 clinical ultrasound scanner and a 15L8 linear array clinical transducer (Siemens Medical Solutions, Mountain View, CA) using the following settings: Cadence Contrast Pulse Sequencing (CPS); 10.5 MHz center frequency; mechanical index, 0.28; dynamic range, 80 dB; gain, –10 (additional details on optimization studies of ultrasound imaging settings are listed in Supplementary Material). A prototype software algorithm (described below) for fast, near real-time ultrasonic molecular imaging signal measurements was developed and interfaced with detected raw contrast and B-mode data streams. Contrast and B-mode images were displayed side-by-side. The transducer was aligned to the center of the tumor, and images were zoomed to a 1.5 cm × 1.5 cm field of view. An acoustic focus zone was placed at the level of the tumor. To decrease potential artifacts from tissue motion, motion compensation software using real-time B-mode tracking (see green dotted box over tumor on Figure 1B and Supplementary Figure S1) of pixel displacement was applied as described recently (27).
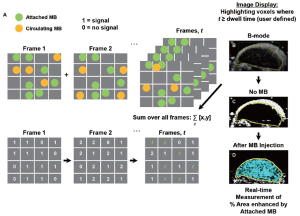
Software algorithm for fast ultrasonic molecular imaging signal measurement
This algorithm was designed to separate imaging signals from freely circulating and molecularly attached microbubbles in near real-time (Figure 1; see also Supplementary Material). Since ultrasonic imaging signals from circulating microbubbles are only transiently measurable whereas signals from attached microbubbles are stationary within a designated sample volume, the principle of the new algorithm was based on the measurement of the “dwell time” of the microbubble signal in the sample volume of the ultrasound transducer (which samples the resolution volume of about 0.15 [x] × 0.2 [y] × 0.8 [z] mm3 (0.024 mm3) near the transmit and geometric focus of the transducer). Measurements of the “dwell time” were accomplished first by recording the presence (encoded as “1”) or absence (encoded as “0”) of contrast imaging signal above system thermal noise (threshold set to 20% of full scale, or 50 out of 255 grey levels; for details see Supplementary Material) in the sample volume with each frame, and adding the values for the entire predefined acquisition time (set at 30 sec). Then, only imaging signals from microbubbles with a “dwell time” of ≥80% of the predefined 30-sec acquisition time (≥24 sec) were recorded as signal derived from attached microbubbles. The threshold of 24 sec was defined from preceding optimization experiments that demonstrated maximum signal from attached microbubbles at minimal signal from freely circulating microbubbles at a ≥80% dwell time (see Supplementary Material).
With this new software algorithm, attached microbubble signal was quantified as percent contrast coverage area, which is the cross sectional area of detected contrast signal relating to the presence of targeted microbubbles within the ROI area (that is, the percentage of voxels with detected contrast signal divided by the total number of voxels within the ROI; see below). An advantage of this new approach is that the cross sectional coverage area is independent of factors which can affect the linearized signal quantified by the traditional approach (see below) such as the attenuation of the ultrasound signal over the imaging path length, which becomes critical when scaling from small animal experiments to humans with known variability in height and weight (as opposed to mice that have similar dimensions) and with different anatomical locations of pathologies compared to the transducer position. Furthermore, the measurement of the cross sectional coverage area is independent of the magnitude of the microbubble non-linear response which depends on the microbubble diameter (which usually varies within a few micrometers) and the resonance frequency of the microbubbles (which depends on the size of the microbubbles). This variability of the linearized signal requires some form of normalization which can be difficult and time consuming in a clinical setting.
Intra-animal comparison between fast and traditional method of ultrasonic molecular imaging signal quantification
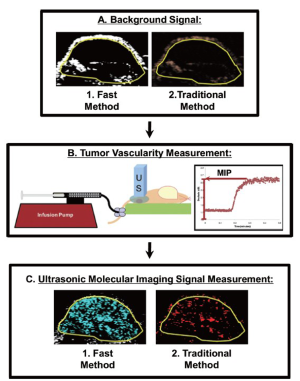
In order to directly compare the new dwell-time based fast ultrasonic molecular imaging quantification approach (henceforth fast method; unit: percent contrast area) with imaging signal measurements using the traditional destruction/replenishment method (6,9,10,28) (henceforth traditional method; unit: linear arbitrary units), data sets using both methods were acquired in all animals during the same imaging session at 7 min after injection of either control or VEGFR2-targeted microbubbles (see Supplementary Material). Data sets using the fast method were collected during 30 sec as described above and saved in DICOM format to the memory of the ultrasound machine. Imaging data sets using the traditional method were collected as described (6,9,10,28). In brief, contrast mode images were acquired for 30 sec, followed by a 3-sec high intensity destruction pulse (acoustic pressure, 5.0 MPa; mechanical index, 1.9) to destroy all microbubbles within the field of view and followed by a post-destruction 30 sec data acquisition. Ultrasonic molecular imaging signal acquired by both techniques was normalized to tumor vascularity measured as described (29,30) (Figure 2); normalization of tumor vascularity was accomplished by taking the near real-time signal (fast method; measured signal at 7 minutes post-injection of microbubbles minus the background signal (no microbubbles; collected first as in Figure 2) divided by the maximum intensity persistence (MIP) imaging signal (measured in linear arbitrary units) that was acquired during the bolus administration of microbubbles and which is a measure of tissue vascularity (29,30).
To assess repeatability of in vivo imaging signal measurements using both methods, a second injection of VEGFR2-targeted MB (with 30 minute waiting period between injections for microbubble clearance as described above) was performed in a subgroup of 6 tumor-bearing mice and data acquisition was performed using both the near real-time and traditional methods as described above.
Monitoring anti-angiogenic therapy
Feasibility of the fast method for monitoring treatment effects following anti-angiogenic therapy was tested in an additional 14 tumor-bearing mice (Figure 3). Baseline (0 h) measurements of ultrasonic molecular imaging signal were obtained in all mice following VEGFR2-targeted microbubble administration. Mice were then randomized into two groups: mice in group 1 (n=7) were treated with an anti-VEGF antibody (B20-4.1.1; Genentech, South San Francisco, CA; cross-reacting with both murine and human VEGF; 10 mg/kg i.p.; volume, 150 μL). Mice in group 2 (n=7) received daily i.p. saline only (150 μL of sterile 0.9% saline; Fisher Scientific, Pittsburgh, PA). Ultrasonic molecular imaging signal measurements normalized to tumor perfusion as described above were performed every 24 hours up to 72 hours in all mice (Figure 3).

Imaging data analysis
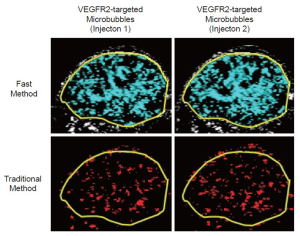
All ROI-based data sets were analyzed in random order by one reader blinded to the type of microbubbles (control vs. VEGFR2-targeted) and to the type of mice (treated vs. non-treated). Data sets acquired using the fast method were quantified as imaging signal (acquired at 7 minutes post-injection of microbubbles) minus the background signal and was displayed in blue (Figures 1, 2). Data sets acquired using the traditional method were post-processed by subtracting post-destruction imaging signal from pre-destruction imaging signal both numerically and by image subtraction using Matlab software (differential pixels were displayed as red; Figure 2) (18).
Ex vivo analysis of VEGFR2 expression
After the 72-hour ultrasound scan, mice were euthanized and human colon cancer xenografts were excised (Figure 3) and frozen in Optimum Cutting Temperature (Tissue-Tek; Fisher Scientific, Pittsburgh, PA). Tumor tissue sections (8 μm; mounted on glass microscope slides) were sliced from the center plane, in close proximity to the field of view used for US scanning. After brief fixation in cold acetone, tissue sections were then double-stained for both VEGFR2 (primary antibody: 1:100 of rabbit anti-mouse VEGFR2 antibody; Cell Signaling; Danvers, MA; secondary antibody: 1:300 of goat anti-rabbit Cy3-conjugated IgG; Jackson Immunoresearch; West Grove, PA) and for the endothelial cell marker CD31 (primary antibody: 1:100 of rat anti-mouse CD31 antibody; BD Pharmingen; San Diego, CA; secondary antibody: 1:300 of Dylight 488 (green)-conjugated goat anti-rat IgG; Jackson Immunoresearch; West Grove, PA). Tissue sections were incubated with primary antibody at 4 ˚C, and 18 hours later, they were washed in phosphate-buffered saline (PBS) and incubated with secondary antibodies for 30 minutes at room temperature. Fluorescent micrographs were obtained at the same exposure using a Zeiss AxioImager DIC/Fluorescence microscope (magnification: 100×) and Zeiss monochrome AxioCam CCD digital camera (Carl Zeiss Light Microscopy; Thornwood, NY).
Statistical analysis
All data were expressed as mean ± standard deviation. The one-sample Wilcoxon rank and t-tests were used for paired comparisons. Repeatability of the two imaging methods was calculated using the interclass correlation coefficient (ICC) including 95% confidence intervals (CI), calculated using the bootstrap method. ICCs were defined as follows: an ICC of 0-0.20 indicated no agreement between two consecutive measurements; an ICC of 0.21-0.40, poor agreement; an ICC of 0.41-0.60, moderate agreement; an ICC of 0.61-0.80, good agreement; and an ICC greater than 0.80, excellent agreement (31). Correlation of contiguous sample measurements were assessed by calculating the Pearson correlation coefficient with the 95% CI calculations based on Fisher’s transformation. A mixed regression model was applied for comparing longitudinal measurements with and without anti-angiogenic therapy. All statistical analyses were computed with R.2.10.1 software (www.r-project.org), and a P-value less than 0.05 was considered statistically significant.
Results
In vivo ultrasonic molecular imaging measurements
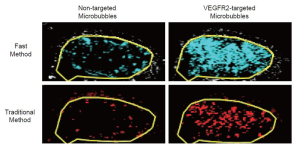
Overall, there was excellent correlation between the fast and the traditional method in terms of in vivo ultrasonic molecular ultrasound imaging signal quantifications following administration of both non-targeted and VEGFR2-targeted microbubbles in human colon cancer xenografts (R2=0.93; P<0.001; 95% CI; 0.91, 0.99). Both the fast (P=0.002) and the traditional method (P=0.03) showed significantly higher imaging signal following administration of VEGFR2-targeted microbubbles compared to control microbubbles, confirming binding specificity of molecularly-targeted microbubbles to VEGFR2 (Figure 4, Table 1).
| Table 1 Ultrasonic molecular imaging signal intensities measured with the new fast and traditional methods following injection of VEGFR2-targeted and non-targeted contrast microbubbles | |||||||
| Imaging signal with non-targeted MB | Imaging signal with VEGFR2-targeted MB (1st Injection ) | Imaging signal with VEGFR2-targeted MB (2nd Injection) | |||||
| Fast method | Traditional method | Fast method | Traditional method | Fast method | Traditional method | ||
| 8.97 ± 4.91 | 1.56 ± 0.78 | 46.46 ± 8.19 | 4.04 ± 1.29 | 47.01 ± 1.53 | 4.08 ± 1.58 | ||
| Note: Numbers are mean ± standard deviation; MB: microbubbles; Units: Fast method, percent contrast area; Traditional method, linearized arbitrary units | |||||||
Reproducibility of in vivo ultrasonic molecular imaging measurements
In the subgroup of tumor-bearing animals that underwent two subsequent VEGFR2-targeted microbubble injections 30 minutes apart, both the fast and the traditional method showed excellent reproducibility of ultrasonic molecular imaging signal measurements. For the fast method, the measured US signal in the tumors was 46.46±18.19% contrast area after the first contrast agent injection and was not significantly different (P=0.82) after the second injection (47.01±21.53% contrast area), indicating excellent agreement (ICC=0.87; 95% CI; 0.69, 0.99). Similarly, for the traditional method, the measured ultrasonic molecular imaging signal in the tumors was 4.04±1.29 a.u. after the first contrast agent injection and was not significantly different (P=0.69) after the second injection (4.08±1.58 a.u.), indicating excellent agreement (ICC=0.87; 95% CI; 0.53, 0.99) (Figure 5). Further analysis of the reproducibility of tumor vascularity measurements in the same tumor-bearing mice confirmed excellent reproducibility of tumor vascularity measurements without statistically significance differences (P=0.23) between repeated measurements (ICC=0.85; 95% CI: 0.71, 0.99).
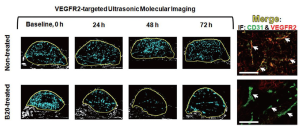
In vivo monitoring of anti-angiogenic therapy in human colon cancer xenografts using fast method of ultrasonic molecular imaging signal quantification
At baseline, tumor volumes in treated (2,174±690 mm3) and non-treated (1,811±1,513 mm3) mice were not significantly different (P=0.31).
Following anti-angiogenic therapy, ultrasonic molecular imaging signal decreased by an average of 41±10% over all time points (P=0.05), while in non-treated animals imaging signal increased by an average of 54±8% (P=0.11). This was confirmed with ex vivo immunofluorescence that showed low level VEGFR2 expression on CD31-positive tumor vessels in treated tumors and high levels of VEGFR2 in non-treated mice (Figure 6).
Tumor volumes, in contrast, significantly increased in both treatment groups (P<0.001) compared to baseline measurements with non-treated tumors growing at significantly faster rate (average increase of tumor volume within 72 hours: 121%; P=0.02) compared to treated tumors (average increase of tumor volume within 72 hours: 36%; P=0.02).
Discussion
Due to the lack of ionizing radiation, widespread availability of ultrasound in medical imaging, and high spatial and temporal resolution, the combination of ultrasound with molecular imaging capabilities has gained great scientific attention in recent years (1-3,6,15). Several preclinical studies have shown that ultrasonic molecular imaging allows highly sensitive detection of molecular markers over-expressed in cancer (8-10,19,26), better monitoring of cancer response to treatment compared to tumor size measurements using morphological-anatomical imaging (2,11-14), and molecular profiling of cancer (8-10) that may eventually help stratifying cancer patients based on biological properties of the tumors. However, to translate this promising imaging approach for improved patient care in the clinic, several technical developments are still needed (3,6,15). While recent research has shown progress towards the design and testing of novel clinical grade targeted ultrasound contrast agents as one of the major prerequisites for clinical translation (26,32), improved imaging data collection and quantification tools that facilitate the workflow of ultrasonic molecular imaging in a clinical environment are critically needed.
In this research, we explored a fast approach of ultrasonic molecular imaging data collection on a clinical ultrasound system to differentiate molecularly-attached contrast microbubbles from freely circulating microbubbles in a human colon cancer xenograft model in vivo. This new approach obviates the need for a separate destructive pulse and time-consuming post-processing, thereby facilitating the work-flow of ultrasonic molecular imaging. In a direct intra-animal comparison in the same mice and during the same imaging sessions, we compared the fast quantitative ultrasonic molecular imaging approach with the traditional destruction/replenishment quantification approach as a reference standard that had been validated in previous studies (10,26,32). Our results showed excellent quantitative correlation between both techniques with significantly higher imaging signal after injection of VEGFR2-targeted microbubbles compared to negative control non-targeted microbubbles. Furthermore, our results suggest that the new fast imaging and quantification approach is highly reproducible with almost identical quantitative values obtained at imaging exams 30 minutes apart.
We also tested the potential of the new imaging approach for longitudinal quantitative monitoring of the molecular effects of anti-angiogenic therapy in human colon cancer xenografts. As early as 24 hours following anti-angiogenic therapy administration, the molecular imaging signal decreased by on average 41% while there was a substantial increase of the imaging signal in non-treated mice. Further ex vivo analysis of VEGFR2 expression levels confirmed decreased receptor levels in treated mice and increased levels in non-treated mice. Our in vivo imaging results are in line with findings of a recent study using the traditional destruction/replenishment approach that also showed substantially decreased ultrasonic molecular imaging signal in a mouse tumor model by on average 41% after 24 hours after anti-angiogenic therapy initiation (14). Overall, our results indicate that the fast ultrasonic molecular imaging approach allows in vivo assessment and monitoring of early anti-angiogenic treatment effects in cancer at the molecular level.
Other approaches to differentiate molecularly attached from freely circulating microbubbles for ultrasonic molecular imaging have recently been explored (16,33-35). Using an avidin-coated cellulose microtube phantom, it has been shown that microtube-attached biotin-containing microbubbles have different spectral characteristics compared to freely circulating microbubbles (16). For attached contrast microbubbles, the fundamental spectral intensity increased by up to 22 dB, the second harmonic component increased, and the difference between the fundamental intensity and the intensity of the 2nd and 3rd harmonic components increased for attached versus freely circulating microbubbles in those experiments (16). However, while this approach could be exploited for real-time differentiation of attached from freely circulating microbubbles (16), in vivo proof of this concept is still missing. Another principle to differentiate attached from free microbubbles using a low-pass filter (7-frame moving average filter) was tested in a biotin-containing gelatin vessel phantom using avidin-containing microbubbles and a dedicated small animal ultrasound system (33). Using subharmonic imaging to first separate background tissue from microbubble signal, this approach allowed separation of bound microbubble imaging signal from free microbubbles in vitro (33). The principle of low-pass filtering to separate bound from free microbubble signal was also shown in another in vitro phantom study using biotin-coated microbubbles targeted to avidin-coated cellulose tubes and a clinical ultrasound system equipped with a transducer that transmits at a low frequency and receipts at a high frequency (35). However, both approaches have only been applied to in vitro phantom studies and in vivo confirmation is still needed (33,35). Furthermore, the avidin-biotin interactions between microbubbles and the tubing used for above-mentioned phantom studies represent one of the strongest chemical bonds not reflecting the binding kinetics of ligands with in vivo receptors. Also the tubing size used for the phantom studies (200 μm) is bigger than the size reported for tumor capillaries [~10-50 μm (36)] which may affect the flow dynamics and attachment properties of microbubbles. In our study, we explore an alternative approach to differentiate attached from freely circulating microbubbles based on a threshold-determined “dwell time” calculation. Our study brings previous studies on differentiation of attached from free microbubbles to several different levels: First, to the best of our knowledge, our study is the first in vivo assessment of a fast near real-time ultrasonic molecular imaging approach in an animal model of cancer. Second, we used a clinical grade VEGFR2-targeted contrast agent that models ligand receptor interactions expected to occur in human cancers and the technique was tested on a clinical ultrasound scanner to simulate a clinical imaging environment. Third, we performed a direct intra-animal comparison of the new approach with the traditional quantification approach for ultrasonic molecular imaging as a reference standard. Fourth, we further validated the new quantification approach for non-invasive in vivo monitoring of anti-angiogenic therapy in a clinically relevant mouse model of human colon cancer treated with the murine correlate of the clinically used anti-angiogenic agent bevacizumab. Finally, we further confirmed our new ultrasonic molecular imaging approach by immunofluorescence analysis of VEGFR2 expression on tumor vascular endothelial cells.
This new quantification technique may help translate ultrasonic molecular imaging into the clinic by enabling rapid visualization and quantification of relative expression levels of molecular imaging targets in vivo. However, we acknowledge the following limitations. First, the new imaging approach was tested for the limited field-of-view of a currently clinically available two-dimensional (2D) ultrasound transducer. Further developments are needed to move from 2D to three-dimensional (3D) quantification using phased array 3D ultrasound transducers to measure the spatial distribution of molecular marker expressions and the overall expression levels within the full extent of the tumors. Second, this imaging approach can only detect imaging signal from attached microbubbles which is above the front-end thermal system noise floor. Furthermore, once an imaging signal from attached microbubbles is registered above the system noise floor threshold, the software cannot determine the number of microbubbles contained within each sample volume (i.e., if the average number of attached microbubble exceeds one microbubble per voxel, the molecular receptor density will be underestimated by a factor equal to the receptor density). However, at a spatial resolution of 0.024 mm3 with the transducer used in our study and given the observation using intravital microscopy that only small amounts of targeted microbubbles are actually retained in vivo, on the order of 10 microbubbles per mm3 (37,38), the predicted average number of attached microbubbles is on the order of one or fewer per voxel in our study. In fact, at densities of on average one or fewer attached microbubbles per voxel, there is a linear relationship between the measured cross sectional area and the number of attached microbubbles which would allow absolute estimations of the molecular receptor density in tissues. This likely explains the excellent correlation (R2=0.93) between the percent contrast coverage area measured with the fast method and the linearized signal measured with the traditional method in our study. However, if binding efficiency of targeted microbubbles can be further improved in the future with increased expected densities of attached microbubbles beyond one microbubble per voxel [for example by combining different strategies such as acoustic radiation force techniques (39) or using microbubbles with increased binding affinity (19,40)], the proposed fast quantification approach may only provide semiquantitative information (though, which will still be helpful to assess relative changes of imaging signal for example during treatment or to perform ultrasonic molecular imaging guided biopsies) (41).
In conclusion, our results suggest that the new ultrasonic molecular imaging approach implemented on a clinical ultrasound system allows accurate and reproducible near real-time quantification and monitoring of VEGFR2 expression levels in human colon cancer xenografts in mice during anti-VEGF therapy. Along with next generation clinical grade molecularly targeted contrast agents as used in this study, the improved work-flow afforded by the new quantification approach may facilitate clinical translation of ultrasonic molecular imaging.
Acknowledgements
This work has been supported by the NIH R21 CA139279 grant, the R01 CA155289-01A1, the SMIS NIH fellowship program NIH/NCI R25 CA11868, and the Canary Foundation. We acknowledge the technical expertise and support by Andrew Olsen, PhD (Neuroscience Microscopy Service) and of Timothy Doyle, PhD (Small Animal Imaging Facility) at Stanford University. We also acknowledge Francois Tranquart, MD PhD (Bracco Suisse SA, Geneva, Switzerland) for providing BR55, and Siemens, Mountain View CA, USA for provision of the clinical ultrasound system.
Disclosure: Ismayil Guracar is an employee of Siemens. All other authors declare no conflict of interest.
References
- Klibanov AL. Microbubble contrast agents: targeted ultrasound imaging and ultrasound-assisted drug-delivery applications. Invest Radiol 2006;41:354-62.
- Leong-Poi H. Molecular imaging using contrast-enhanced ultrasound: evaluation of angiogenesis and cell therapy. Cardiovasc Res 2009;84:190-200.
- Hwang M, Lyshchik A, Fleischer AC. Molecular sonography with targeted microbubbles: current investigations and potential applications. Ultrasound Q 2010;26:75-82.
- Wilson SR, Burns PN. Microbubble-enhanced US in body imaging: what role? Radiology 2010;257:24-39.
- Deshpande N, Pysz MA, Willmann JK. Molecular ultrasound assessment of tumor angiogenesis. Angiogenesis 2010;13:175-88.
- Pysz MA, Willmann JK. Targeted contrast-enhanced ultrasound: an emerging technology in abdominal and pelvic imaging. Gastroenterology 2011;140:785-90.
- Kircher MF, Willmann JK. Molecular Body Imaging: MR Imaging, CT, and US. Part I. Principles. Radiology 2012;263:633-43.
- Sanna V, Pintus G, Bandiera P, et al. Development of polymeric microbubbles targeted to prostate-specific membrane antigen as prototype of novel ultrasound contrast agents. Mol Pharm 2011;8:748-57.
- Bzyl J, Lederle W, Rix A, et al. Molecular and functional ultrasound imaging in differently aggressive breast cancer xenografts using two novel ultrasound contrast agents (BR55 and BR38). Eur Radiol 2011;21:1988-95.
- Deshpande N, Ren Y, Foygel K, et al. Tumor angiogenic marker expression levels during tumor growth: longitudinal assessment with molecularly targeted microbubbles and US imaging. Radiology 2011;258:804-11.
- Korpanty G, Carbon JG, Grayburn PA, et al. Monitoring response to anticancer therapy by targeting microbubbles to tumor vasculature. Clin Cancer Res 2007;13:323-30.
- Palmowski M, Huppert J, Ladewig G, et al. Molecular profiling of angiogenesis with targeted ultrasound imaging: early assessment of antiangiogenic therapy effects. Mol Cancer Ther 2008;7:101-9.
- Palmowski M, Peschke P, Huppert J, et al. Molecular ultrasound imaging of early vascular response in prostate tumors irradiated with carbon ions. Neoplasia 2009;11:856-63.
- Pysz MA, Foygel K, Rosenberg J, et al. Antiangiogenic cancer therapy: monitoring with molecular US and a clinically translatable contrast agent (BR55). Radiology 2010;256:519-27.
- Deshpande N, Needles A, Willmann JK. Molecular ultrasound imaging: current status and future directions. Clin Radiol 2010;65:567-81.
- Zhao S, Kruse DE, Ferrara KW, et al. Acoustic response from adherent targeted contrast agents. J Acoust Soc Am 2006;120:EL63-9.
- Ferrara K, Pollard R, Borden M. Ultrasound microbubble contrast agents: fundamentals and application to gene and drug delivery. Annu Rev Biomed Eng 2007;9:415-47.
- Willmann JK, Paulmurugan R, Chen K, et al. US imaging of tumor angiogenesis with microbubbles targeted to vascular endothelial growth factor receptor type 2 in mice. Radiology 2008;246:508-18.
- Willmann JK, Kimura RH, Deshpande N, et al. Targeted contrast-enhanced ultrasound imaging of tumor angiogenesis with contrast microbubbles conjugated to integrin-binding knottin peptides. J Nucl Med 2010;51:433-40.
- Deshpande N, Lutz AM, Ren Y, et al. Quantification and monitoring of inflammation in murine inflammatory bowel disease with targeted contrast-enhanced US. Radiology 2012;262:172-80.
- Sonne C, Xie F, Lof J, et al. Differences in definity and optison microbubble destruction rates at a similar mechanical index with different real-time perfusion systems. J Am Soc Echocardiogr 2003;16:1178-85.
- Grishenkov D, Kari L, Brodin LK, et al. In vitro contrast-enhanced ultrasound measurements of capillary microcirculation: comparison between polymer- and phospholipid-shelled microbubbles. Ultrasonics 2011;51:40-8.
- Sciallero C, Paradossi G, Trucco A. A preliminary in vitro assessment of polymer-shelled microbubbles in contrast-enhanced ultrasound imaging. Ultrasonics 2012;52:456-64.
- Miller DL, Averkiou MA, Brayman AA, et al. Bioeffects considerations for diagnostic ultrasound contrast agents. J Ultrasound Med 2008;27:611-32.
- Caruso G, Valentino B, Salvaggio G, et al. Ultrastructural biologic effects of sonography with pulse inversion and microbubble contrast in rabbit liver. J Clin Ultrasound 2005;33:106-11.
- Pochon S, Tardy I, Bussat P, et al. BR55: a lipopeptide-based VEGFR2-targeted ultrasound contrast agent for molecular imaging of angiogenesis. Invest Radiol 2010;45:89-95.
- Pysz MA, Guracar IM, Foygel K, et al. Quantitative assessment of tumor angiogenesis using real-time motion-compensated contrast-enhanced ultrasound imaging. Angiogenesis 2012. Epub ahead of print.
- Anderson CR, Hu X, Zhang H, et al. Ultrasound molecular imaging of tumor angiogenesis with an integrin targeted microbubble contrast agent. Invest Radiol 2011;46:215-24.
- Palmowski M, Lederle W, Gaetjens J, et al. Comparison of conventional time-intensity curves vs. maximum intensity over time for post-processing of dynamic contrast-enhanced ultrasound. Eur J Radiol 2010;75:e149-53.
- Pysz MA, Foygel K, Panje CM, et al. Assessment and monitoring tumor vascularity with contrast-enhanced ultrasound maximum intensity persistence imaging. Invest Radiol 2011;46:187-95.
- Faria JR, Aarao AR, Jimenez LM, et al. Inter-rater concordance study of the PASI (Psoriasis Area and Severity Index). An Bras Dermatol 2010;85:625-9.
- Anderson CR, Rychak JJ, Backer M, et al. scVEGF microbubble ultrasound contrast agents: a novel probe for ultrasound molecular imaging of tumor angiogenesis. Invest Radiol 2010;45:579-85.
- Needles A, Couture O, Foster FS. A method for differentiating targeted microbubbles in real time using subharmonic micro-ultrasound and interframe filtering. Ultrasound Med Biol 2009;35:1564-73.
- Patil AV, Rychak JJ, Allen JS, et al. Dual frequency method for simultaneous translation and real-time imaging of ultrasound contrast agents within large blood vessels. Ultrasound Med Biol 2009;35:2021-30.
- Hu X, Zheng H, Kruse DE, et al. A sensitive TLRH targeted imaging technique for ultrasonic molecular imaging. IEEE Trans Ultrason Ferroelectr Freq Control 2010;57:305-16.
- Ryschich E, Schmidt E, Maksan SM, et al. Expansion of endothelial surface by an increase of vessel diameter during tumor angiogenesis in experimental and hepatocellular and pancreatic cancer. World J Gastroenterol 2004;10:3171-4.
- Leong-Poi H, Christiansen J, Klibanov AL, et al. Noninvasive assessment of angiogenesis by ultrasound and microbubbles targeted to alpha(v)-integrins. Circulation 2003;107:455-60.
- Rychak JJ, Klibanov AL, Hossack JA. Acoustic radiation force enhances targeted delivery of ultrasound contrast microbubbles: in vitro verification. IEEE Trans Ultrason Ferroelectr Freq Control 2005;52:421-33.
- Gessner RC, Streeter JE, Kothadia R, et al. An in vivo validation of the application of acoustic radiation force to enhance the diagnostic utility of molecular imaging using 3-d ultrasound. Ultrasound Med Biol 2012;38:651-60.
- Willmann JK, Lutz AM, Paulmurugan R, et al. Dual-targeted contrast agent for US assessment of tumor angiogenesis in vivo. Radiology 2008;248:936-44.
- Kaneko OF, Willmann JK. Ultrasound for molecular imaging and therapy in cancer. Quant Imaging Med Surg 2012;2:81-91.
Supplementary methods
Optimization of contrast-enhanced ultrasound imaging settings
The contrast-enhanced ultrasound imaging parameters were adjusted to maximize the signal intensity from attached microbubbles and to minimize imaging signal from system noise (Figure S1). This was accomplished by testing different threshold levels and measuring the imaging signal within the tumor region of interest (ROI). At a gain of -10 the system noise was just barely visible in the CPS contrast imaging mode and was used for subsequent experiments. At a threshold setting of 50 (20% of full scale), system noise was nearly completely rejected while signals from microbubbles are mostly retained. All ultrasound imaging parameters and settings are summarized in Table S1. In all animals, tissue equalization (TEQ) depth-gain levels were aligned to the same levels according to the line displayed (Figure S1; TEQ, red arrow).
| Table S1 Contrast-enhanced ultrasound imaging parameters and settings | |
| Center Frequency | 10.5 MHz |
| Mechanical Index | 0.28 |
| Dynamic Range | 80 dB |
| Threshold Signal Intensity (noise floor reject) | 50 (Scale 0 (black) → 255 (white)) |
| Gain | -10 |
| Dwell Time | 80% (Percent of total Acquisition Time) |
| Acquisition Time | 30 seconds |
| Waiting Time for Targeted Microbubbles to Attach | 7 minutes dB, decibels; MHz, mega Hertz. User-defined dwell time was designated as |
| dB, decibels; MHz, mega Hertz. User-defined dwell time was designated as a percentage of the total time. In this study, it was set to 80%. For the set acquisition time of 30 seconds, this means that signal that was recorded for 24 seconds (80% of 30 seconds) was counted as targeted contrast signal from attached microbubbles | |
Optimization of the waiting time for targeted microbubble attachment
The optimal time to wait before acquiring attached microbubble signal was evaluated by analyzing the wash-out rates of non-targeted and targeted microbubbles in an additional n=5 LS174T tumor-bearing mice. Time-intensity curves in real-time linear intensity CPS mode were collected for 10 minutes during the intravenous injection [via tail vein catheter using an infusion pump (Genie Plus; Kent Scientific, Torrington, CT) set at a constant rate of 1.2 mL/min] of 5×107 non-targeted, control microbubbles and VEGFR2-targeted microbubbles. A waiting period of at least 30 minutes under continuous destruction (mechanical index, 1.9) was used to clear microbubbles in circulation between injection of the two microbubble types (control and VEGFR2-targeted), and background levels were collected to ensure that microbubbles were cleared. Time intensity curve data sets were exported to and graphed in Microsoft Excel and the time at which the curves for VEGFR2-targeted and control microbubbles separated in parallel was recorded as the optimal time for microbubbles to attach while background signal from freely circulating microbubbles is minimal. This average time was 6.6±0.6 minutes (range, 5.8-7.4 minutes); therefore, a waiting time of 7 minutes to allow microbubbles to attach was chosen for subsequent experiments.
Optimization of dwell time
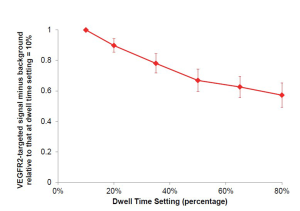
The optimal dwell time (defined as a percentage of the set acquisition time of 30 seconds) was evaluated in an additional n=4 LS174T tumor-bearing mice using the near real-time software. After background signal (no MBs) was collected, VEGFR2-targeted microbubbles were injected intravenously [as described in manuscript (Figure 2)], and ultrasound molecular imaging signal was collected 7 minutes post-injection with the fast method and background subtraction. Various dwell times (percentages) were tested: 10%, 20%, 35%, 50%, 65%, and 80%; with a 30 second acquisition time, this corresponded to dwell times of 3, 6, 10.5, 15, 19.5, and 24 seconds. Percent contrast area of attached VEGFR2-targeted microbubbles was calculated as signal acquired at 7 min post-injection minus background signal (no MBs), and compared relatively to other dwell percentages by normalizing to dwell percentage of 10% (by dividing the calculated result of each signal (fast method signal post-injection of microbubbles minus the background signal (no microbubbles)), by the signal obtained at dwell percentages of 10%; Supplementary Figure S2). Minimal differences were observed in the VEGFR2-targeted microbubble signal obtained with dwell times ranging between 50% and 80%. For subsequent experiments a dwell time of 80% of the 30-sec acquisition time (24 sec) was used.