
Mycobacterial cavity on chest computed tomography: clinical implications and deep learning-based automatic detection with quantification
Introduction
Mycobacterium is a genus of aerobic acid-fast bacteria that cause diverse diseases to humans. The Mycobacterium genus comprises more than 150 species and includes important human pathogens such as Mycobacterium tuberculosis and nontuberculous mycobacteria (NTM), with representatives including the Mycobacterium avium complex, Mycobacterium kansasii, and Mycobacterium abscessus complex (1-3). Pulmonary tuberculosis (TB) is one of the top 10 causes of mortality, and 7.0 million new cases were notified worldwide in 2018 (4). TB remains a major global health problem as it is easily transmitted from human to human by droplet nuclei (5). Positive sputum smear test for acid-fast bacilli indicates the capability of disease transmission (6). On the other hand, NTM are ubiquitous environmental pathogens in the soil and water, without person-to-person transmission (7). The frequency of NTM isolation among mycobacterial isolates (8) and the prevalence of NTM pulmonary disease (NTM-PD) have been increasing worldwide (9) and in Korea (10). Not all the patients diagnosed as NTM-PD are coerced into treatment, and clinicians traditionally determined patients who need to be medicated (11).
A cavity is a radiologic and pathologic hallmark of pulmonary mycobacterial disease and one of the most relevant features to disease burden and prognosis (12-15). Each cavity contains an enormous burden of bacilli, resulting in high infectivity, drug resistance, and treatment failure in pulmonary TB (16,17). Similarly, a cavity in NTM-PD is a risk factor for reduced sputum culture conversion and mortality (9,15), and cavity-associated biofilms may explain the incurability of M. abscessus disease (18). Particularly, the NTM treatment guideline in 2020 suggests treatment initiation for cavitary NTM at diagnosis rather than watchful waiting followed by treatment initiation when disease progressed (11).
Computed tomography (CT) is a crucial imaging modality for evaluating lung parenchymal lesions in pulmonary TB and NTM-PD, including cavities (5,9,19). Assessing the cavity burden on CT images from patients with pulmonary TB and NTM-PD may help monitor the treatment response of drug-resistant TB and the treatment initiation of NTM-PD requiring long-term treatment. Nevertheless, it can be cumbersome to measure the baseline cavity burden and to track changes in the burden on CT images, particularly for volumetric measurements that require substantial time and human resources. Deep learning algorithms are increasingly used to automatically detect and quantify pulmonary diseases on CT images and replace the laborious efforts of radiologists. Deep learning models were developed for various lung diseases on chest CT such as lung cancer screening and diffuse lung disease classification (20). Several models were developed for mycobacterial diseases, for example, to differentiate NTM from Mycobacterium tuberculosis (21), to diagnose pulmonary TB (22), or to evaluate the activity or severity pulmonary TB (23,24). Yet, to our knowledge, there has been no previous model to detect and quantify the CT cavity itself, which is a hallmark of mycobacterial pulmonary disease. Among numerous deep learning methods, convolutional neural network (CNN), represented by U-Net, is an effective model that consists of multiple layers to find the most significant features from the hierarchical information (25-27). In particular, nnU-Net automatically configures the network by itself without manual parameter optimization and provides better segmentation performance than a conventional U-Net (28).
Herein, we hypothesized that (I) CT cavity volume was associated with sputum positivity in TB and the necessity of treatment in NTM-PD and (II) deep learning could automatically detect and quantify mycobacterial cavities on chest CT. We aimed (I) to investigate the association of CT cavity volume with sputum smear positivity in TB and necessity of treatment in NTM-PD, and (II) to develop a three-dimensional (3D) nnU-Net model for the automatic detection and volumetric quantification of mycobacterial cavities on CT images. We present the following article in accordance with the STARD reporting checklist (available at https://qims.amegroups.com/article/view/10.21037/qims-22-620/rc).
Methods
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The institutional review board of Seoul National University Hospital approved this study involving no more than minimal risk to the subjects and waived the requirement for informed consent.
Patient population and baseline characteristics
A study coordinator retrospectively searched for the records of consecutive patients who were treated for pulmonary TB and NTM-PD and underwent chest CT scans between November 2011 and July 2019 in a single tertiary hospital. We applied the following eligibility criteria: (I) microbiological confirmation of TB on sputum smear or culture, or polymerase chain reaction confirmations from respiratory specimens or NTM-PD diagnosis according to 2007 American Thoracic Society and Infectious Diseases Society of America guideline (11); and (II) available thin-section CT images at diagnosis. Two radiologists (JH Hong and I Yoon) reviewed the CT images of the patients in consensus to determine the presence of a cavity in the lung window setting (window width, 1,500 HU; level, −700 HU), resulting in a total of 108 and 93 patients with cavitary TB and NTM-PD, respectively. We then introduced a group of respectively 98 and 93 age- and sex-matched patients with non-cavitary TB and NTM-PD among a large pool of non-cavitary patients as non-cavitary mycobacterial diseases were more prevalent than cavitary diseases in the hospital. Finally, a total of 392 patients were included in this study (Figure 1): 206 TB patients (mean age, 57±18 years; male-to-female ratio, 152:54) and 186 NTM-PD patients (mean age, 64±9.7 years; male-to-female ratio, 64:122).
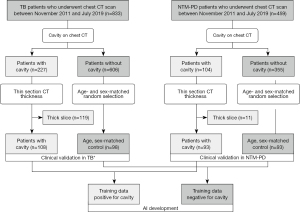
The coordinator collected clinical information, including the treatment initiation due to clinical or radiologic disease aggravation in NTM-PD patients by reviewing the electronic medical record. The treatment initiation was determined as decision to treat by duty physician during follow-up period in the included patients between 2011–2019 based on the 2007 guideline (11). We chose NTM treatment initiation to examine the clinical implication of CT cavity volume as the 2020 NTM treatment guideline suggests treatment initiation for cavitary diseases (11), while the former 2007 guideline did not (11). The degrees of sputum smear positivity were based on the hospital’s clinical practice and defined as followings; negative (no bacilli detected on smear), equivocal positive (one to two bacilli found on 300 microscopic fields), 1+ (one to nine bacilli found per 100 fields), 2+ (one to nine bacilli found per 10 fields), 3+ (one to nine bacilli per field), and 4+ (more than nine bacilli found per field).
Among the 392 mycobacterial pulmonary disease patients included in this study, 242 were never-smokers and the remaining 150 were ever-smokers. The average time interval between the sputum smear test and the CT scan of TB patients was 29±58 days. For NTM-PD cases who were treated, the mean interval between treatment initiation and the CT scan was 124±276 days. Furthermore, out of a total of 206 TB patients enrolled in our study, 19 patients had previous treatment history and 187 were newly diagnosed at the time of our study. The drug susceptibility test results of 206 TB cases consisted of 166 drug-susceptible, 16 multidrug-resistant, one extensively drug-resistant, two other poly-drug resistant cases and 21 cases without drug sensitivity test results. The baseline characteristics of the study population are presented in Table 1.
Table 1
| Characteristic | All (n=392) | TB cohort | NTM cohort | |||
|---|---|---|---|---|---|---|
| Cavity (−) (n=98) | Cavity (+) (n=108) | Cavity (−) (n=93) | Cavity (+) (n=93) | |||
| Age† (years) | 60±15 | 57±18 | 56±18 | 63±9.7 | 64±9.7 | |
| Sex | ||||||
| Male | 216 | 73 | 79 | 32 | 32 | |
| Female | 176 | 25 | 29 | 61 | 61 | |
| Smoking history | ||||||
| Never-smoker | 242 | 53 | 47 | 74 | 68 | |
| Ex-smoker | 67 | 13 | 16 | 17 | 21 | |
| Current smoker | 83 | 32 | 45 | 2 | 4 | |
| Symptom | ||||||
| Cough | 50% (195/392) | 51% (50/98) | 56% (61/108) | 43% (40/93) | 47% (44/93) | |
| Sputum | 53% (209/392) | 37% (36/98) | 45% (49/108) | 71% (66/93) | 62% (58/93) | |
| Hemoptysis | 15% (58/392) | 5% (5/98) | 14% (15/108) | 16% (15/93) | 25% (23/93) | |
| Weight loss | 12% (47/392) | 9% (9/98) | 18% (19/108) | 6% (6/93) | 14% (13/93) | |
| Sputum positivity | 29% (57/196) | 14% (14/98) | 44% (43/98) | Not applicable | Not applicable | |
| Treatment initiation | 51% (94/186) | Not applicable | Not applicable | 22% (20/93) | 80% (74/93) | |
| Cavity | ||||||
| Mean number†‡ | 2.3±2.1 | 0 | 2.6±2.4 | 0 | 2.1±1.8 | |
| Mean volume (cm3)†‡ | 13.8±36.7 | 0 | 11.3±14.6 | 0 | 16.4±50.6 | |
| Lobar location | ||||||
| RUL | 179 | 0 | 110 | 0 | 69 | |
| RML | 24 | 0 | 6 | 0 | 18 | |
| RLL | 105 | 0 | 43 | 0 | 62 | |
| LUL | 117 | 0 | 79 | 0 | 38 | |
| LLL | 62 | 0 | 38 | 0 | 24 | |
†, the values are presented as means ± standard deviation; ‡, considering only the cavity positive cases. NTM, nontuberculous mycobacteria; TB, tuberculosis; RUL, right upper lobe; RML, right middle lobe; RLL, right lower lobe; LUL, left upper lobe; LLL, left lower lobe.
CT acquisition
The total number of chest CT scans was 400, with 194 from NTM-PD patients and 206 from pulmonary TB patients. Eight NTM-PD patients with cavities had two CT scans, and the rest of the patients had one CT scan.
All CT scans were performed using one of the following 16- or higher-channel multi-detector CT scanner of several manufacturers: Siemens Healthineers [Somatom Definition (n=73), Sensation 16 (n=53), Somatom Force (n=24), Somatom Definition Flash (n=10) Somatom Scope (n=2), Sensation 64 (n=1), and Somatom Spirit (n=1)]; Philips Medical Systems [Brilliance 64 (n=117), Brilliance iCT 256 (n=35), Ingenuity CT (n=15), and IQon Spectral CT (n=11)]; Toshiba [Aquilion One (n=42), Asteion (n=1)]; GE Healthcare [Discovery CT750 HD (n=6), Lightspeed VCT (n=2), Lightspeed Ultra (n=1), Brightspeed (n=1), Hispeed (n=1), Optima CT660 (n=1), and Revolution CT (n=1)]; Shimadzu equipment (SCT-7000TS (n=2)]. CT scans were obtained in the supine position at full inspiration with (n=106) or without (n=294) contrast enhancement. The CT tube voltage and current settings were 100–120 kVp and a standard-dose (n=137) or low-dose (n=263) scan with automatic exposure control. The field of view of the CT scans covered a whole thorax (median value of 319 mm × 319 mm, ranging from 238 mm × 238 mm to 420 mm × 420 mm) and the matrix sizes were uniformly 512×512. CT images were reconstructed mostly with a slice thickness of 1 mm (n=357) or 1.25 mm (n=5) and a minor number of cases with larger thickness only used for clinical feature assessment and not for deep learning model development (2 mm, n=3; 2.5 mm, n=4; 3 mm, n=31). A sharp reconstruction kernel was applied to the CT images.
Segmentation and volumetric measurement of cavities
CT images were uploaded and displayed in a lung window setting in commercially available segmentation software (MEDIP PRO v2.0.0.0, MEDICALIP Co. Ltd.). A cavity was defined as an abnormal air-filled space in the lung parenchyma with perceptible wall within pulmonary consolidation, a mass, or a nodule based on the Fleischner Society glossary (29). All cavities on CT images were initially segmented by a third-year radiology resident (I Yoon) in every axial CT image slice using a semi-automatic method, producing volumetric cavity masks. A chest radiologist (SH Yoon, with 15 years of clinical experience in thoracic imaging) reviewed and confirmed the cavity masks. The two radiologists were blinded to the clinical information of the patients. The volume of each segmented mask was calculated in the software.
Model development
Radiologist-derived masks served as reference masks. To build the 3D nnU-Net, we used 298 cases, with 5,862 positive and negative slices each. The positive slices were provided by all CT images containing cavity masks of 199 patients with 1- to 1.25-mm CT images amongst 211 patients who have cavities. In contrast, negative slices were randomly selected from CT images not containing cavity masks from the 199 patients above and additional CT images from eighty patients with 1- to 1.25-mm CT scan amongst 191 patients without cavitary diseases. We assigned 240 cases for training, 20 for tuning, 38 randomly chosen age- and sex-matched cases, including 19 cases with cavities and 19 cases without cavities, were assigned for internal validation.
nnU-Net is an up-to-date deep-learning segmentation scheme that is characterized by automatic configuration of the entire segmentation process regardless of various dataset properties (28). We implemented the nnU-Net codes publicly available on GitHub (https://github.com/MIC-DKFZ/nnUNet). The 3D nnU-Net received an input size of 96×160×160 and used five encoders and five decoders. Every step consists of two convolution modules, each followed by instance normalization and Leaky Rectified Linear Unit (LReLU). The encoder adopted 3×3×3 convolution pooling, whereas the decoder adopted 2×2×2 transposed convolution to restore the feature. Softmax function was used at the final layer and stochastic gradient descent algorithm were used train the network (Figure 2).
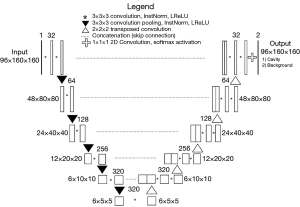
The hyper-parameters, hardware and software used for the model development are listed in Tables 2,3. The simplified code for the model can be found on GitHub (https://github.com/josephnw/cavity-nnunet-pytorch).
Table 2
| Parameter | Value |
|---|---|
| Network initialization | Kaiming He Initialization |
| Initial feature map size | 32 |
| Patch size | 96×160×160 |
| Batch size | 2 |
| Number of convolution operations | 28 |
| Data augmentation | Image scaling, rotation, flip, elastic deformation and gamma transformation |
| Activation function | Leaky ReLU with negative slope of 0.01 |
| Loss function | Dice loss + cross-entropy loss |
| Optimizer | Stochastic gradient descent with Nesterov momentum of 0.99 |
| Learning rate | Polynomial learning rate with initial value of 0.01 and decay scheduler with a power of 0.9 |
| Number of epochs | 1,000 |
ReLU, Rectified Linear Unit.
Table 3
| Parameter | Value |
|---|---|
| CPU | Intel® CoreTM i9-10980XE CPU @ 3.00 GHz ×36 |
| GPU | NVIDIA RTX3090 (24 GB) |
| RAM | 8×32 GB |
| CUDA version | 11.1 |
| Programming language | Python 3.8.10 |
| Deep learning framework | PyTorch 1.11.0, torchvision 0.12.0 |
CPU, central processing unit; GPU, graphics processing unit; RAM, random-access memory; CUDA, compute unified device architecture.
Statistical analysis
Association of CT cavity volume with sputum positivity in TB and treatment necessity for NTM-PD
The Pearson chi-square test was performed to evaluate the correlations between symptoms and the presence of a cavity. The per-case cavity volumes of the TB and NTM-PD patients were compared using the independent sample t-test and the F-test. The independent t-test was performed to compare cavity volumes depending on sputum positivity in TB and the necessity of treatment in NTM-PD. The Spearman correlation coefficient was used to examine the relationship between the degree of sputum smear positivity (on a scale from 0 to 4+) and cavity volume in TB. For the eight NTM-PD patients whose two CT scans on different dates were incorporated in the study, only the CT scan temporally closer to the treatment initiation date was selected. Receiver operating characteristic (ROC) curves were drawn to determine the area under the curve (AUC) for assessing the diagnostic accuracy of CT cavity volume for sputum positivity and necessity of treatment in TB and NTM-PD, respectively. We calculated accuracy measures for optimal cutoff values on the basis of the ROC analysis by maximizing the Youden index (Youden index = sensitivity + specificity −1). The AUC values of the male and female subgroups were compared in TB and NTM-PD, respectively.
Performance evaluation of the 3D nnU-Net model for cavity detection and segmentation on CT images
The per-patient and per-lesion sensitivity, specificity, and accuracy were calculated to assess the ability of the 3D nnU-Net model to detect cavities. Per-patient calculations were carried out utilizing the original 3D nnU-Net detection results of each case. Per-patient sensitivity was determined as the number of cases that had any correctly detected cavities divided by the total number of cavity-positive cases. Per-lesion sensitivity was defined as the number of correctly detected cavities divided by the number of total reference cavities. The false-positive rate was also calculated by dividing the number of false-positive cavities by the total number of cases.
The intraclass correlation coefficient was calculated and Bland-Altman analysis was conducted between the reference cavity volume and the U-Net-derived volume to analyze the consistency of volume measurements made using the two different methods. The paired t-test was performed to compare the mean values of the reference and U-Net-detected cavity volumes. The Dice similarity coefficient, sensitivity, and precision were calculated for a quantitative measurement of the overlap between reference cavities and U-Net-derived masks. Analysis on association of radiologist-detected and 3D nnU-Net-derived CT cavity volume with a clinical index of mycobacterial diseases was described in Appendix 1.
Evaluation for potential clinical utility of the 3D nnU-Net model
Two chest radiologists with 15 and 5 years of clinical experience (SH Yoon, JH Hong) participated in the reader study. First, the two radiologists read the 38 chest CT images of validation cases without the deep learning output to detect and measure the 3D diameters of the CT cavities. Each cavity volume was then estimated by the formula of ellipsoid sphere volume, multiplying 4/3 and π by three perpendicular radii and summated to obtain the cavity volume of each case. Second, the readers reviewed the CT images overlaid with the nnU-Net-driven cavity masks and modified the inappropriate masks in the MEDIP program if necessary.
The intraclass correlation coefficient was calculated, and Bland-Altman analysis was conducted as follows: (I) between the reference cavity volume and the volume measured with or without deep learning to analyze the agreement between the reference volume and measured volume; (II) between the measured volume values of the two readers to test the consistency of volume measurements between them. The paired t-test was performed to compare the mean elapsed time for volume measurement according to the availability of deep learning assistance in each reader.
All statistical analyses were performed using commercially available statistical software [MedCalc v19.1 (RRID:SCR_015044); SPSS v25 (RRID:SCR_002865)].
Results
Association of CT cavity volume and symptom
Among the 392 mycobacterial pulmonary disease patients included in this study, about half of the patients presented with cough (50%) and sputum (53%). Hemoptysis (18.9% vs. 10.5%, P=0.019) and weight loss (15.9% vs. 7.9%, P=0.014) were significantly more frequent in patients with CT cavities than in patients without CT cavities (Table 1).
Cavity characteristics
The number of radiologist-segmented cavities was 276 in the 206 TB cases, including 108 cavity-positive cases; 211 in the 194 NTM-PD cases (from 186 patients including eight with two CTs of different dates), including 101 cavity-positive cases; and 487 in all 400 cases, including 209 cavity-positive cases. The NTM-PD and TB patients had similar mean per-case cavity numbers. The mean number of cavities per chest CT scan of only the cases with cavities was 2.6±2.4 in TB patients, 2.1±1.8 in NTM-PD patients, and 2.3±2.1 in total. The mean volume of cavities regarding only the lesion-positive cases was 11.3±14.6 cm3 in TB patients, 16.4±50.6 cm3 in NTM-PD patients, and 13.8±36.7 cm3 in total. The cavities of NTM-PD cases had a similar volume (P=0.420) but a larger volume range than those of TB cases (P=0.007). The location of the cavities showed a slight predominance for the upper lobes (179 in the right upper lobe, 24 in the right middle lobe, 105 in the right lower lobe, 117 in the left upper lobe, and 62 in the left lower lobe).
Association of CT cavity volume with sputum smear positivity in TB and treatment necessity for NTM-PD
Among the 196 TB patients, 57 had sputum smear-positive test results, and 139 had smear-negative tests. The average per-patient cavity volume was 11 cm3 in sputum smear-positive TB patients and 4.0 cm3 in smear-negative patients (P<0.001). In the 186 NTM-PD patients, the per-patient volume average values were 14 cm3 in the 94 patients with treatment initiation due to clinical or radiologic aggravation and 1.6 cm3 in the 92 patients who did not receive treatment (P=0.020). The degree of smear positivity and cavity volume in TB patients showed a moderate positive relationship (Spearman correlation coefficient, 0.374; P<0.001).
The per-case CT cavity volume provided an AUC of 0.701 [95% confidence interval (CI): 0.620–0.782; P<0.001] with optimal cutoff value of 1.4 cm3 [sensitivity, 0.72 (95% CI: 0.59–0.83); specificity, 0.68 (95% CI: 0.59–0.75)] for sputum positivity in TB and 0.834 (95% CI: 0.773–0.894; P<0.001) with optimal cutoff value of 0.75 cm3 [sensitivity, 0.71 (95% CI: 0.61–0.80); specificity, 0.88 (95% CI: 0.80–0.94)] for treatment necessity of NTM-PD, respectively. The AUCs of the males [0.734 (95% CI: 0.654–0.803) in TB; 0.803 (95% CI: 0.684–0.892) in NTM-PD] and females [0.814 (95% CI: 0.444–0.729) in TB; 0.852 (95% CI: 0.776–0.910) in NTM-PD] did not significantly differ regarding the per-case CT volume and TB sputum positivity (P=0.1779; difference between areas, 0.141; difference 95% CI: −0.0643 to 0.347) and NTM-PD treatment necessity (P=0.4337; difference between areas, 0.0491; difference 95% CI: −0.0739 to 0.172) (Figure 3).
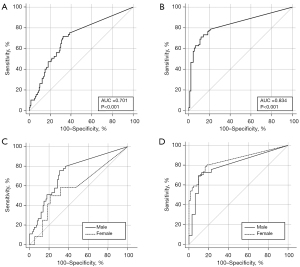
Ability of the 3D nnU-Net model to detect cavities
Forty-nine reference cavities were provided by the radiologists from 19 internal validation cases with cavities. The trained 3D nnU-Net independently detected and segmented cavities on CT images from the same validation cases within seconds, producing volumetric cavity masks. Per-lesion and per-patient comparisons were made between the volume of reference cavity and 3D nnU-Net detected cavities (Tables 4-6).
Table 4
| Cases with or without cavities | Cases with reference cavity | Cases without reference cavity | Sum | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Total | NTM-PD | TB | Total | NTM-PD | TB | Total | NTM-PD | TB | |||
| Cases with 3D nnU-Net-detected cavity | 19 | 9 | 10 | 2 | 1 | 1 | 21 | 10 | 11 | ||
| Cases without 3D nnU-Net-detected cavity | 0 | 0 | 0 | 17 | 8 | 9 | 17 | 8 | 9 | ||
| Sum | 19 | 9 | 10 | 19 | 9 | 10 | 38 | 18 | 20 | ||
3D, three-dimensional; NTM-PD, nontuberculous mycobacteria pulmonary disease; TB, tuberculosis.
Table 5
| Cavity lesions | Reference cavities | No reference cavity | Sum | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Total | NTM-PD | TB | Total | NTM-PD | TB | Total | NTM-PD | TB | |||
| 3D nnU-Net-detected cavities (+) | 41 | 15 | 26 | 18 | 10 | 8 | 59 | 25 | 34 | ||
| 3D nnU-Net-detected cavities (−) | 8 | 3 | 5 | 17 | 8 | 9 | 25 | 11 | 14 | ||
| Sum | 49 | 18 | 31 | 35 | 18 | 17 | 84 | 36 | 48 | ||
3D, three-dimensional; NTM-PD, nontuberculous mycobacteria pulmonary disease; TB, tuberculosis.
Table 6
| Variables | Values |
|---|---|
| Per-case sensitivity† | 1.0 (19/19; 95% CI: 0.8 to 1.0) |
| Per-case specificity‡ | 0.89 (17/19; 95% CI: 0.67 to 0.98) |
| Per-case accuracy§ | 0.95 (36/38; 95% CI: 0.82 to 0.99) |
| Per-lesion sensitivity¶ | 0.84 (41/49; 95% CI: 0.71 to 0.92) |
| False positive ratio†† | 0.47 (18/38) |
†, sensitivity = total number of true positive cases/total number of positive cases; ‡, specificity = total number of true negative cases/total number of negative cases; §, accuracy = number of true positive and true negative cases/total number of cases; ¶, per-lesion based sensitivities = number of true positive lesions/total number of reference lesions; ††, false-positive detections per CT scan = total number of false positive lesions/total number of cases. CI, confidence interval; CT, computed tomography; 3D, three-dimensional.
In the per-lesion analysis, the 3D nnU-Net detected 84 positive lesions in 19 internal validation cases and 19 control cases. Forty-one lesions matched the reference cavities, resulting in per-lesion sensitivity of 0.84 (41/49; 95% CI: 0.71 to 0.92). The 3D nnU-Net model detected 18 false-positive lesions, resulting in a false-positive ratio of 0.47. Fifteen of the 18 false-positive lesions were from all 19 cavity-positive internal validation cases, and the remaining three were from two control cases.
In a per-patient analysis, the 3D nnU-Net gave a positive result in 21 cases, including all 19 internal validation cases and 2 of the 19 cavity-negative cases. The per-patient sensitivity, specificity, and accuracy were 1.0 (19/19; 95% CI: 0.8 to 1.0), 0.89 (17/19; 95% CI: 0.67 to 0.98), and 0.95 (36/38; 95% CI: 0.82 to 0.99), respectively.
Cavity quantification by the 3D nnU-Net model
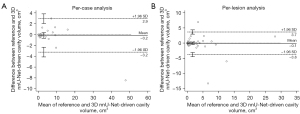
The average per-patient volume from the reference cavities (3.5±7.7 cm3) and that from the 3D nnU-Net-predicted results (3.7±8.9 cm3) were not significantly different (P=0.527). The average per-lesion volumes were 1.6±3.7 cm3 for the reference cavities and 1.7±3.9 cm3 for the valid manually separated masks of the 3D nnU-Net-predicted cavities (P=0.723). The per-patient and per-lesion intraclass correlation coefficients between the reference volume and the U-Net-driven volume were excellent, with values of 0.991 (95% CI: 0.983–0.995) and 0.933 (95% CI: 0.897–0.957), respectively (Table 7). The Bland-Altman plots representing the differences and mean values of the reference cavity volume and the U-Net-derived volume of validation cases are shown in Figure 4. In the per-patient analysis, the mean difference and 95% limits of agreement were −0.16 cm3 and −3.2 (95% CI: −4.1 to −2.4 cm3) to 3.0 cm3 (95% CI: 2.0 to 3.8 cm3), respectively; in the per-lesion analysis, the mean difference and the 95% limits of agreement were −0.07 cm3 and −3.8 (95% CI: −4.5 to −3.1 cm3) to 3.7 cm3 (95% CI: 2.9 to 4.4 cm3), respectively.
Table 7
| Analysis basis | Average volumes of cavities (cm3)† | P values‡ | Intraclass correlation coefficients (95% CI)§ | |
|---|---|---|---|---|
| Reference cavities | 3D nnU-Net-predicted cavities | |||
| Per-patient | 3.5±7.7 | 3.7±8.9 | 0.527 | 0.991 (0.983–0.995) |
| Per-lesion | 1.6±3.7 | 1.7±3.9 | 0.723 | 0.933 (0.897–0.957) |
†, the values are presented as means ± standard deviation; ‡, P values calculated by paired t-test comparing the mean values of the reference and U-Net-detected cavity volumes; §, intraclass correlation coefficient calculated between the reference cavity volume and the U-Net-derived volume. CI, confidence interval; 3D, three-dimensional.

Regarding segmentation accuracy between the manually segmented cavities and the nnU-Net-derived cavities, the mean Dice similarity coefficient was 78.9±10.7, the sensitivity was 81.0±13.5, and the precision was 79.2±13.0 (Figures 5,6).
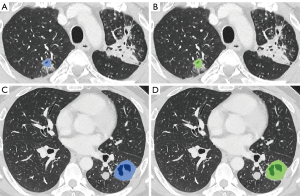
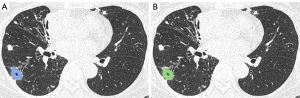
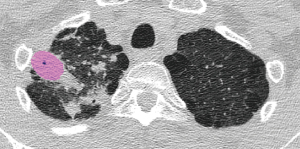
The median per-lesion volume of the false-positive lesions was 0.26 cm3 (mean, 1.0±3.1 cm3). The false-positive lesions were generally small, and 10 of the 18 false-positive lesions had predicted volumes smaller than 0.30 cm3. On the contrary, only 7 of the 41 true-positive lesions exhibited predicted volumes smaller than 0.30 cm3 (Figure 7).
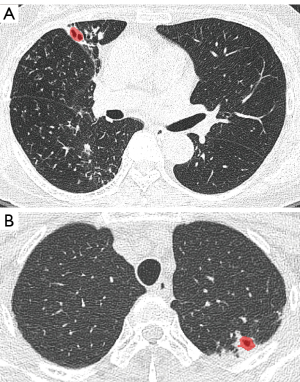
There were eight false negative lesions out of 84 nnU-Net detected cavity lesions. The mean volume of the false negative cavity lesions was 1.9±2.3 cm3 (Figure 8).
Evaluation for potential clinical utility of the 3D nnU-Net model
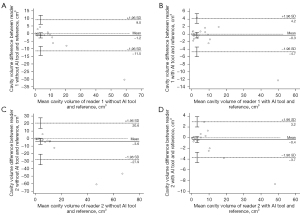
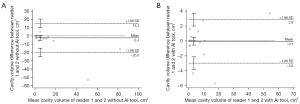
The agreement of the per-case cavity volume with the reference cavities was also greater with deep learning assistance. The intraclass correlation coefficients between the reference cavity volume and reader-assessed cavity volumes increased when 3D nnU-Net-driven cavity masks were provided (0.982 in reader 1 with 95% CI of 0.966–0.991 and 0.990 in reader 2 with 95% CI of 0.980–0.995) compared to absence of the masks (0.934 in reader 1 with 95% CI of 0.872–0.966 and 0.778 in reader 2 with 95% CI of 0.573–0.885). The Bland-Altman plots representing the differences and mean values of the reference cavity volume and volume measured by readers with or without AI assists are shown in Figure 9.
The intraclass correlation coefficient between the per-case cavity volumes measured by two readers was higher in assist of the deep learning model (0.994 with 95% CI of 0.988–0.997) than working only by radiologists themselves (0.916 with 95% CI of 0.839–0956). The Bland-Altman plots visualizing the differences and mean values of the two readers with or without the deep learning outputs are in Figure 10.
The time required for CT cavity assessment was significantly shorter (P<0.001 in reader 1 and P=0.004 in reader 2) with the aid of 3D nnU-Net in both readers (18,568±65,173 s in reader 1 and 2,943±5,737 s in reader 2) than without its assistance (252±179 s in reader 1 and 71±73 s in reader 2).
Discussion
TB cavity volume was associated with smear positivity and the grade of positivity. This finding is in accord with those of previous studies reporting associations of smear positivity with cavitary TB (16,30). The presence of a cavity was the sole risk factor for smear positivity, although some of the smear-negative patients also had cavities on CT images. The cavity volume showed modest diagnostic accuracy for smear positivity (AUC, 0.701) and a moderate linear relationship with bacilli burden (31). This result indicates that the cavity volume reflects the disease burden of TB. Considering that drug-resistant TB is frequently accompanied by cavities and requires biomarker analyses to monitor long-term treatment efficacy, TB cavity volume may be an early surrogate marker for treatment success (32).
Determining treatment initiation is a challenging task in NTM-PD. Clinical risk factors for treatment initiation include symptomatic presentations of night sweats or weight loss, multiple positive results on sputum acid-fast bacteria smears, and co-infection with Aspergillus species (33-35). Chest imaging is mandatory in patients suspected of having NTM-PD, and cavitary disease on radiologic examinations has been found to be associated with treatment initiation (36). Our study showed a similar result, as cavity volume had an AUC of 0.834, highlighting the importance of quantifying cavity volume on CT images in patients with NTM-PD. Furthermore, quantification of the cavity volume on CT may facilitate the better monitoring of treatment response when combined with clinical assessments (33,34,37).
The 3D nnU-Net model showed an excellent diagnostic capability for mycobacterial cavities (sensitivity of 0.84–1.0, specificity of 0.89 and accuracy of 0.95). A few false-positive lesions were found, but most frequently resulted from bronchiectasis (13/18, 72%), followed by non-cavitary nodules (3/18, 17%) and focal consolidation (2/18, 11%). These false-positive lesions were relatively small and readily recognizable by radiologists (Figure 7). Most of the false negative lesions had atypical morphology that seemed to be related to mild disease severity, such as two lesions with small sizes less than 0.1 cm3 and four lesions with tiny central gas-filled spaces (Figure 8). Anyhow further model training with various mycobacterial cavities would keep improving the performance in detecting cavities with non-typical shapes. The U-Net-derived masks of the cavities matched human-driven reference masks with a relatively high mean Dice similarity coefficient of 78.9. The nnU-Net-driven cavity volume also well correlated with the reference volume (intraclass correlation coefficients, 0.933–0.991). When two radiologists assessed the mycobacterial CT cavities, the speed and accuracy were both better with the assistance of deep learning. This implies the potential benefit of our 3D nnU-Net model for the mycobacterial cavity assessment in clinical practice. The current model will assist in quantitative radiologic research on mycobacterial diseases and other cavitary pulmonary lesions with a similar CT shape (e.g., cavitary lung cancer) by reducing the time and effort needed for segmentation.
Several limitations exist in this study. First, although this study analyzed a consecutive group of patients with NTM and TB, the patients were collected from a single tertiary hospital, and the number of patients was relatively small. Second, we did not examine the performance of the 3D nnU-Net model in an external validation dataset. Nevertheless, the diverse distribution of CT scanners in the training data may have contributed to the network’s performance. Third, the preparation of cavity masks might be arbitrary in some cases abutting consolidation or the chest wall, as the cavity margin was not well separated from the adjacent structures. Fourth, NTM treatment was initiated based on clinical or radiologic aggravation rather than a single established indication. This may reflect the difficulty of clinical practice in managing NTM-PD patients. Fifth, all the cavities were assumed to be caused by mycobacterial pulmonary disease and the possibility of cavity formation by other etiologies such as aspergillosis was not concerned. Sixth, the CT acquisition parameters were heterogeneous due to the usage of varying CT scanners and both non-enhanced and enhanced CT images to increase CT training datasets as much as possible.
In conclusion, CT cavity volume was associated with sputum positivity and the necessity of treatment in TB and NTM-PD, respectively. The 3D nnU-Net model could automatically detect mycobacterial cavities and quantified the cavity volume on chest CT images. Intractable mycobacterial diseases, including drug-resistant TB and NTM-PD, often accompany cavities but treatment monitoring tools are scarce. Given that quantifying cavitary volumes is almost impractical by human readers due to substantial demands for time and human resources, our model may serve as a practical tool for assessing disease severity and treatment response by quantifying baseline cavity volumes and tracking its volumetric changes during treatment in cavitary mycobacterial diseases.
Acknowledgments
Funding: This work was supported by the Korea Medical Device Development Fund grant funded by the Korea government (the Ministry of Science and ICT, the Ministry of Trade Industry and Energy, the Ministry of Health & Welfare, Republic of Korea, the Ministry of Food and Drug Safety) (No. 202011A03).
Footnote
Reporting Checklist: The authors have completed the STARD reporting checklist. Available at https://qims.amegroups.com/article/view/10.21037/qims-22-620/rc
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://qims.amegroups.com/article/view/10.21037/qims-22-620/coif). SHY joined the MEDICAL IP, Co., Ltd. As a chief medical officer since November 2020. JNW is an employer and machine learning researcher of MEDICAL IP, Co., Ltd. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The institutional review board of Seoul National University Hospital approved this study involving no more than minimal risk to the subjects and waived the requirement for informed consent.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- King HC, Khera-Butler T, James P, et al. Environmental reservoirs of pathogenic mycobacteria across the Ethiopian biogeographical landscape. PLoS One 2017;12:e0173811. [Crossref] [PubMed]
- Primm TP, Lucero CA, Falkinham JO 3rd. Health impacts of environmental mycobacteria. Clin Microbiol Rev 2004;17:98-106. [Crossref] [PubMed]
- Rastogi N, Legrand E, Sola C. The mycobacteria: an introduction to nomenclature and pathogenesis. Rev Sci Tech 2001;20:21-54. [Crossref] [PubMed]
- Global tuberculosis report 2019. World Health Organization, 2019.
- Jeong YJ, Lee KS. Pulmonary tuberculosis: up-to-date imaging and management. AJR Am J Roentgenol 2008;191:834-44. [Crossref] [PubMed]
- Shaw JB, Wynn-Williams N. Infectivity of pulmonary tuberculosis in relation to sputum status. Am Rev Tuberc 1954;69:724-32. [PubMed]
- Thomson RM, Yew WW. When and how to treat pulmonary non-tuberculous mycobacterial diseases. Respirology 2009;14:12-26. [Crossref] [PubMed]
- Koh WJ, Chang B, Jeong BH, et al. Increasing Recovery of Nontuberculous Mycobacteria from Respiratory Specimens over a 10-Year Period in a Tertiary Referral Hospital in South Korea. Tuberc Respir Dis (Seoul) 2013;75:199-204. [Crossref] [PubMed]
- Stout JE, Koh WJ, Yew WW. Update on pulmonary disease due to non-tuberculous mycobacteria. Int J Infect Dis 2016;45:123-34. [Crossref] [PubMed]
- Jeon D. Infection Source and Epidemiology of Nontuberculous Mycobacterial Lung Disease. Tuberc Respir Dis (Seoul) 2019;82:94-101. [Crossref] [PubMed]
- Daley CL, Iaccarino JM, Lange C, et al. Treatment of Nontuberculous Mycobacterial Pulmonary Disease: An Official ATS/ERS/ESCMID/IDSA Clinical Practice Guideline. Clin Infect Dis 2020;71:e1-e36. [Crossref] [PubMed]
- Gomes M, Saad Júnior R, Stirbulov R. Pulmonary tuberculosis: relationship between sputum bacilloscopy and radiological lesions. Rev Inst Med Trop Sao Paulo 2003;45:275-81. [Crossref] [PubMed]
- Rathman G, Sillah J, Hill PC, et al. Clinical and radiological presentation of 340 adults with smear-positive tuberculosis in The Gambia. Int J Tuberc Lung Dis 2003;7:942-7. [PubMed]
- Matsuoka S, Uchiyama K, Shima H, et al. Relationship between CT findings of pulmonary tuberculosis and the number of acid-fast bacilli on sputum smears. Clin Imaging 2004;28:119-23. [Crossref] [PubMed]
- Ito Y, Hirai T, Maekawa K, et al. Predictors of 5-year mortality in pulmonary Mycobacterium avium-intracellulare complex disease. Int J Tuberc Lung Dis 2012;16:408-14. [Crossref] [PubMed]
- Palaci M, Dietze R, Hadad DJ, et al. Cavitary disease and quantitative sputum bacillary load in cases of pulmonary tuberculosis. J Clin Microbiol 2007;45:4064-6. [Crossref] [PubMed]
- Ong CW, Elkington PT, Friedland JS. Tuberculosis, pulmonary cavitation, and matrix metalloproteinases. Am J Respir Crit Care Med 2014;190:9-18. [Crossref] [PubMed]
- Fennelly KP, Ojano-Dirain C, Yang Q, et al. Biofilm Formation by Mycobacterium abscessus in a Lung Cavity. Am J Respir Crit Care Med 2016;193:692-3. [Crossref] [PubMed]
- Nachiappan AC, Rahbar K, Shi X, et al. Pulmonary Tuberculosis: Role of Radiology in Diagnosis and Management. Radiographics 2017;37:52-72. [Crossref] [PubMed]
- Hwang EJ, Park CM. Clinical Implementation of Deep Learning in Thoracic Radiology: Potential Applications and Challenges. Korean J Radiol 2020;21:511-25. [Crossref] [PubMed]
- Wang L, Ding W, Mo Y, et al. Distinguishing nontuberculous mycobacteria from Mycobacterium tuberculosis lung disease from CT images using a deep learning framework. Eur J Nucl Med Mol Imaging 2021;48:4293-306. [Crossref] [PubMed]
- Li X, Zhou Y, Du P, Lang G, Xu M, Wu W. A deep learning system that generates quantitative CT reports for diagnosing pulmonary Tuberculosis. Applied Intelligence 2020;51:4082-93. [Crossref]
- Ma L, Wang Y, Guo L, Zhang Y, Wang P, Pei X, Qian L, Jaeger S, Ke X, Yin X, Lure FYM. Developing and verifying automatic detection of active pulmonary tuberculosis from multi-slice spiral CT images based on deep learning. J Xray Sci Technol 2020;28:939-51. [Crossref] [PubMed]
- Yan C, Wang L, Lin J, et al. A fully automatic artificial intelligence-based CT image analysis system for accurate detection, diagnosis, and quantitative severity evaluation of pulmonary tuberculosis. Eur Radiol 2022;32:2188-99. [Crossref] [PubMed]
- Lee JG, Jun S, Cho YW, et al. Deep Learning in Medical Imaging: General Overview. Korean J Radiol 2017;18:570-84. [Crossref] [PubMed]
- Zhou T, Tan T, Pan X, et al. Fully automatic deep learning trained on limited data for carotid artery segmentation from large image volumes. Quant Imaging Med Surg 2021;11:67-83. [Crossref] [PubMed]
- Li K, Liu K, Zhong Y, et al. Assessing the predictive accuracy of lung cancer, metastases, and benign lesions using an artificial intelligence-driven computer aided diagnosis system. Quant Imaging Med Surg 2021;11:3629-42. [Crossref] [PubMed]
- Isensee F, Jaeger PF, Kohl SAA, et al. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 2021;18:203-11. [Crossref] [PubMed]
- Hansell DM, Bankier AA, MacMahon H, et al. Fleischner Society: glossary of terms for thoracic imaging. Radiology 2008;246:697-722. [Crossref] [PubMed]
- Ors F, Deniz O, Bozlar U, et al. High-resolution CT findings in patients with pulmonary tuberculosis: correlation with the degree of smear positivity. J Thorac Imaging 2007;22:154-9. [Crossref] [PubMed]
- Ko JM, Park HJ, Kim CH, et al. The relation between CT findings and sputum microbiology studies in active pulmonary tuberculosis. Eur J Radiol 2015;84:2339-44. [Crossref] [PubMed]
- Chung MJ, Lee KS, Koh WJ, et al. Drug-sensitive tuberculosis, multidrug-resistant tuberculosis, and nontuberculous mycobacterial pulmonary disease in nonAIDS adults: comparisons of thin-section CT findings. Eur Radiol 2006;16:1934-41. [Crossref] [PubMed]
- Rawson TM, Abbara A, Kranzer K, et al. Factors which influence treatment initiation for pulmonary non-tuberculous mycobacterium infection in HIV negative patients; a multicentre observational study. Respir Med 2016;120:101-8. [Crossref] [PubMed]
- Tatem G, Jaffery H, Digiovine B, Betensley AD. Characteristics Associated With Initiation of Treatment for Nontuberculous Mycobacterial (NTM) Pulmonary Disease in the Non-HIV Population. Chest 2003;124:96S. [Crossref]
- Provoost J, Valour F, Gamondes D, et al. A retrospective study of factors associated with treatment decision for nontuberculous mycobacterial lung disease in adults without altered systemic immunity. BMC Infect Dis 2018;18:659. [Crossref] [PubMed]
- Haworth CS, Banks J, Capstick T, et al. British Thoracic Society guidelines for the management of non-tuberculous mycobacterial pulmonary disease (NTM-PD). Thorax 2017;72:ii1-ii64. [Crossref] [PubMed]
- Oshitani Y, Kitada S, Edahiro R, et al. Characteristic chest CT findings for progressive cavities in Mycobacterium avium complex pulmonary disease: a retrospective cohort study. Respir Res 2020;21:10. [Crossref] [PubMed]


